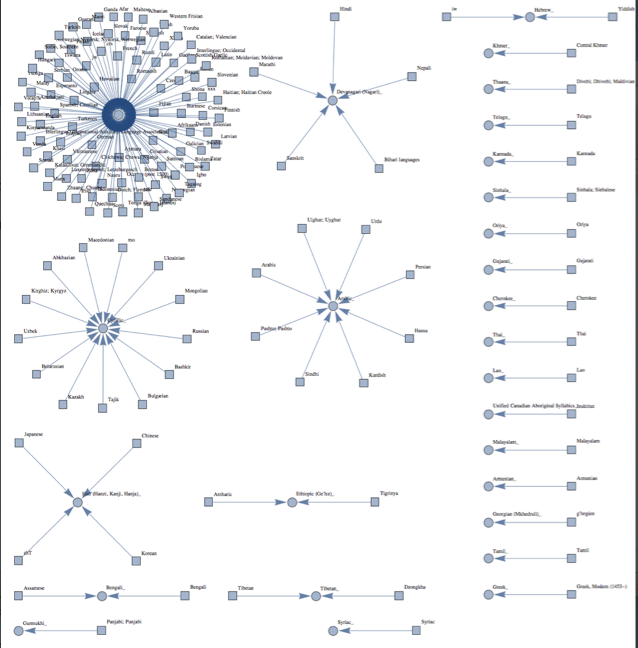

Full image
Notebook

Install["/Users/Yaroslav/mathematica/Pythonika"];
SetDirectory[NotebookDirectory[]];
Needs["Bulatov`sdp2`"];
theta = lovaszTheta@GraphData[{"Paley", 13}];
RootApproximant@theta

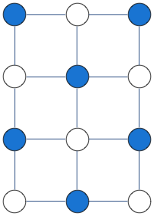
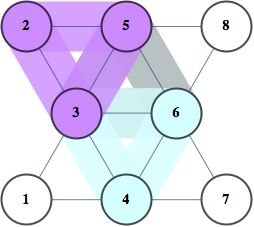
g = GridGraph[{4, 3}];
A = AdjacencyMatrix@g;
n = Length@VertexList@g;
u = ConstantArray[1, n];
vars = x /@ Range@n;
cons = And @@ (-1 <= x[#] <= 1 & /@ Range@n);
solution=vars /. Last[Maximize[{-vars.A.vars, cons}, vars, Integers]]Needs["Bulatov`sdp1`"];
Install["/Users/Yaroslav/mathematica/Pythonika"];
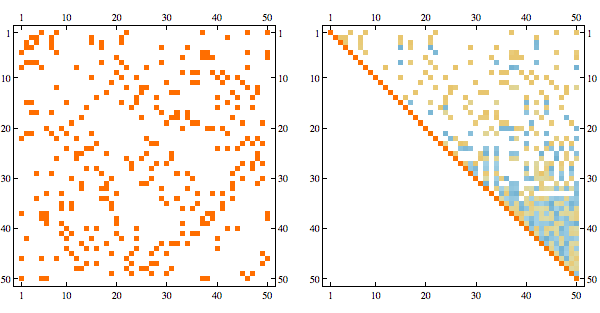
gram = solveDualSDP[-A, IdentityMatrix[n], u];
solution = gramRound[gram];

t = AffineTransform[{{{-(1/Sqrt[2]), -(1/Sqrt[6]),
1/Sqrt[3]}, {1/Sqrt[2], -(1/Sqrt[6]), 1/Sqrt[3]}, {0, Sqrt[2/3],
1/Sqrt[3]}}, {1/3, 1/3, 1/3}}];
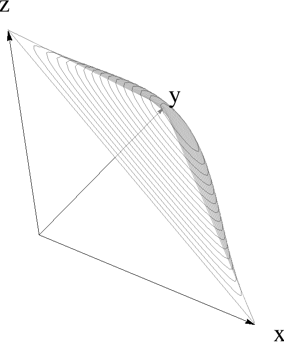
graphics = simplexPlot[5 Sin[#1 #2 #3] &, Plot3D];
shape = Cases[graphics, _GraphicsComplex];
gr = Graphics3D[{Opacity[.5], GeometricTransformation[shape, t]},
Axes -> False, Boxed -> False, Lighting -> "Neutral"];
Show[gr, axes[1, 1, 1, 0.05, 0.02]]words = ReadList["challenge2input.txt", Word];
mat = Map[Characters, words] /. {"O" -> 0, "X" -> 1};
inf[i_] := (inf[i] =
1 + If[Total[mat[[i]]] == 0, 0,
Total[inf /@ Flatten@Position[mat[[i]], 1]]]);
influences = inf /@ Range@Length@mat - 1;
top3 = (Reverse@Sort@influences)[[;; 3]];
answer1 = StringJoin[ToString /@ top3]
mintasks[total_] := (
types = {2, 3, 17, 23, 42, 98};
vars = x /@ Range@Length@types;
poscons = And @@ (# >= 0 & /@ vars);
First@Minimize[{Total[vars], vars.types == total && poscons}, vars,
Integers]
);
counts = mintasks /@ {2349, 2102, 2001, 1747};
answer2 = Times @@ counts

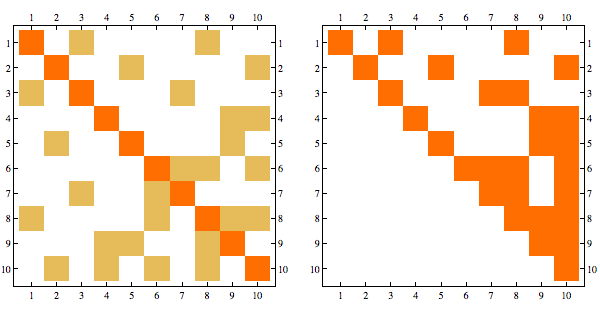
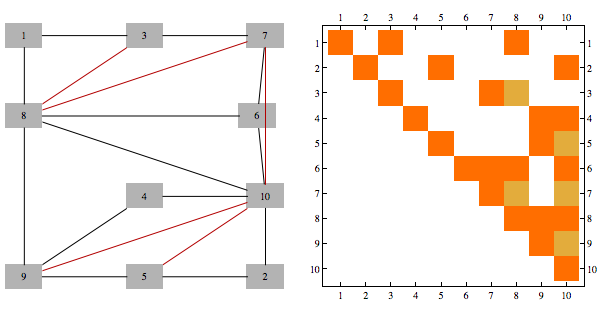
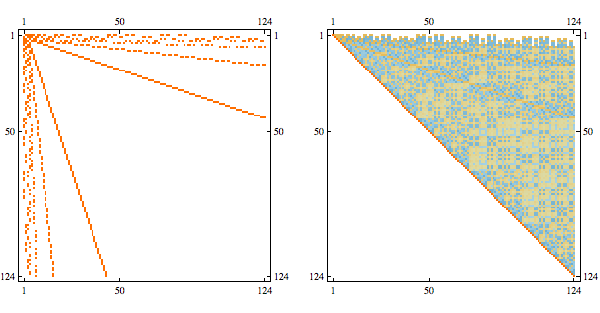
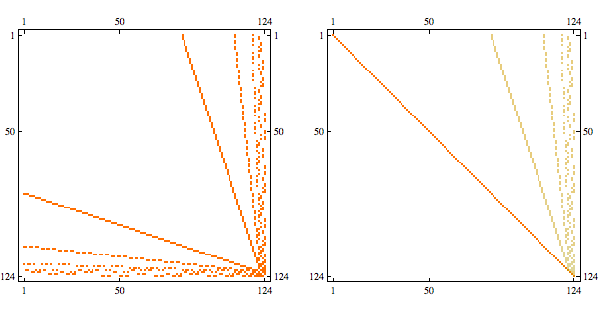
Needs{"Bulatov`chordal`"];
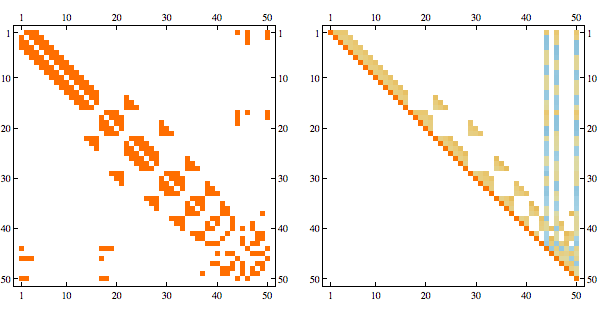
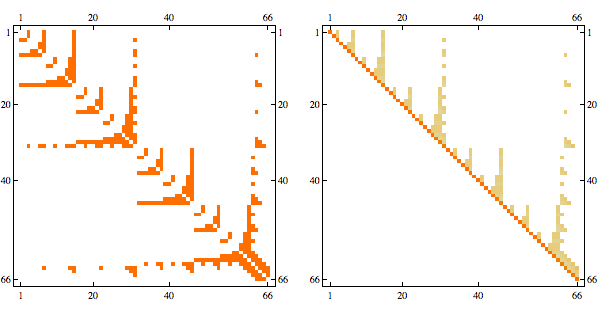
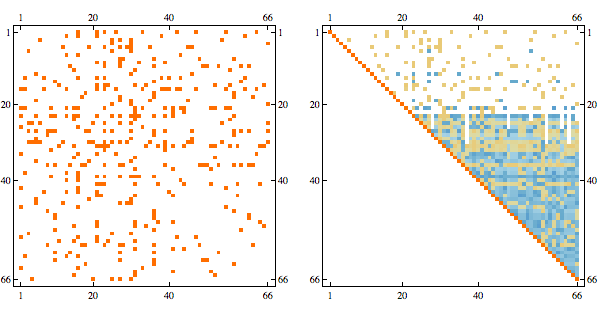
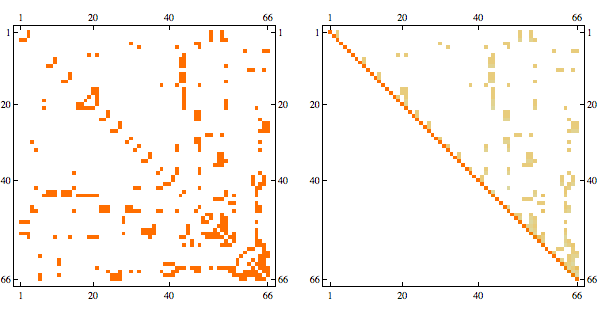
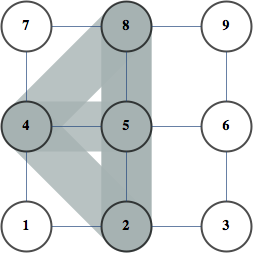
order = getMinFillOrder[AdjacencyGraph@Unitize@A];
CholeskyDecomposition[A[[Reverse@order, Reverse@order]]]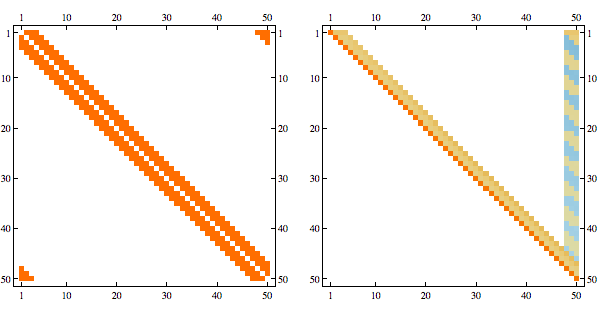

 Notebook+packages
Notebook+packages
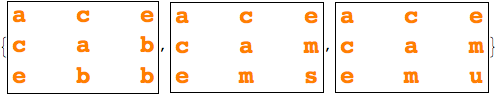
 Notebook
Notebook
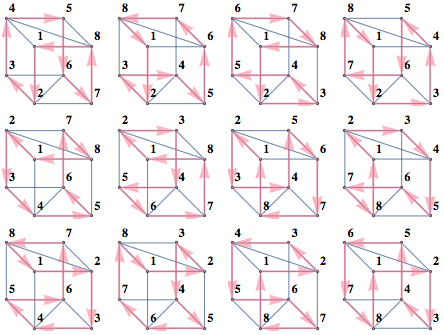
sats = SatisfiabilityInstances[formula, vars, 20];
drawInstance /@ sats
decodeInstance /@ sats
Needs["Bulatov`showGraphs`"];
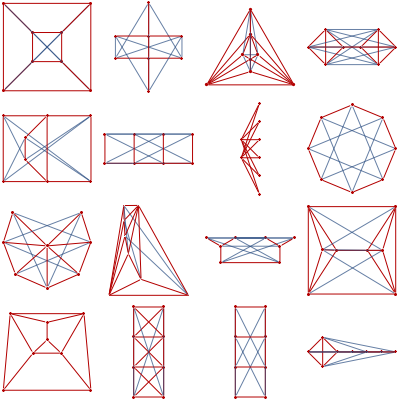
showGraphs[12, "Connected", ! "Tree", "Planar", gridSize -> 6]

 Notebook+package
Notebook+package
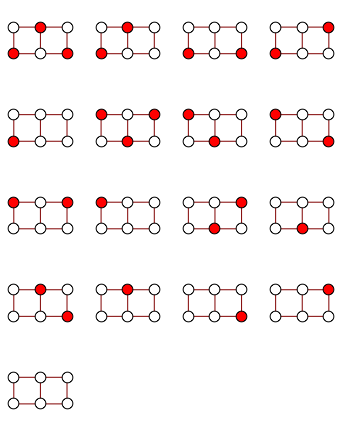
gg = GridGraph[{2, 3}];
edge2clause[e_] := ! (e[[1]] && e[[2]]);
clauses = edge2clause /@ EdgeList[gg];
formula = And @@ clauses;
vars = VertexList[gg];
instances = SatisfiabilityInstances[formula, vars, 1000];
independentSets = Extract[vars, Position[#, True]] & /@ instances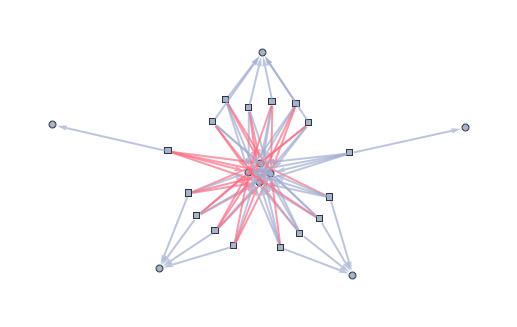
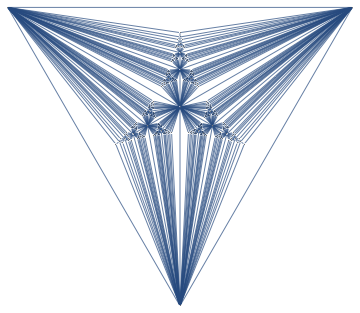
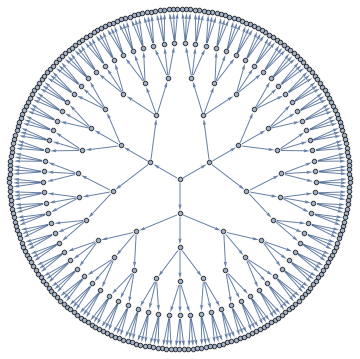
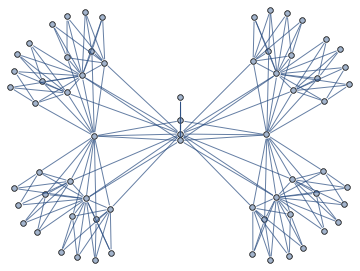
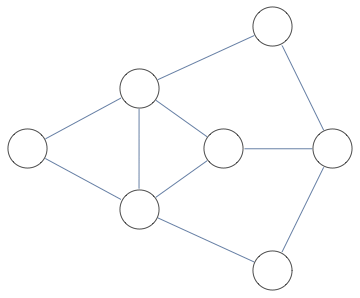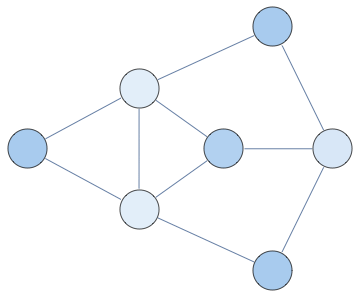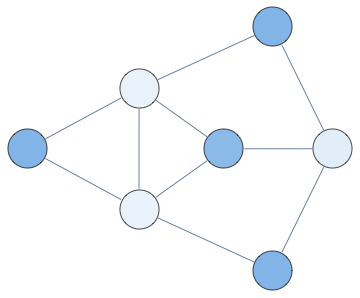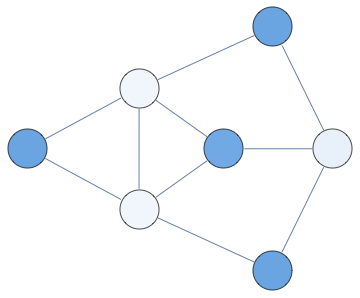
<< Bulatov`treeDecomposition`;
g = GraphData[{"Apollonian", 5}];
{nodes, edges} = findTreeDecomposition[g];
GraphPlot[Rule @@@ edges]
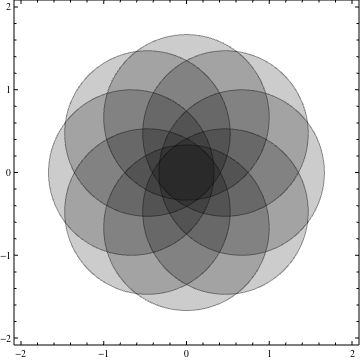
nc = 8;
ineqs = Table[(x - 2/3 Cos[i 2 Pi/nc])^2 + (y -
2/3 Sin[i 2 Pi/nc])^2 < 1, {i, 0, nc - 1}];
Show[Table[
RegionPlot[ineqs[[k]], {x, -2, 2}, {y, -2, 2}, PlotPoints -> 35,
PlotStyle -> Opacity[.2]], {k, 1, nc}]]
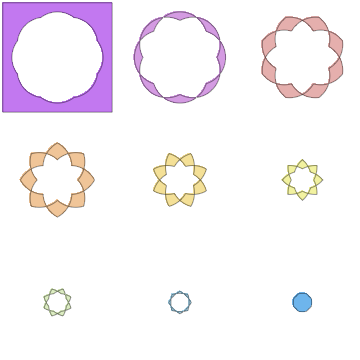
plots = Table[
RegionPlot[
BooleanCountingFunction[{k}, ineqs], {x, -2, 2}, {y, -2, 2},
PlotPoints -> 100, Frame -> None,
PlotStyle -> ColorData["Pastel"][k/nc]], {k, 0, nc}];
GraphicsGrid@Partition[plots, 3]
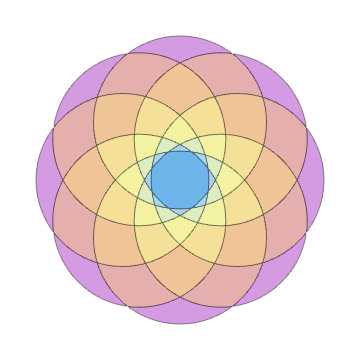
Show[Rest[plots]]
 | 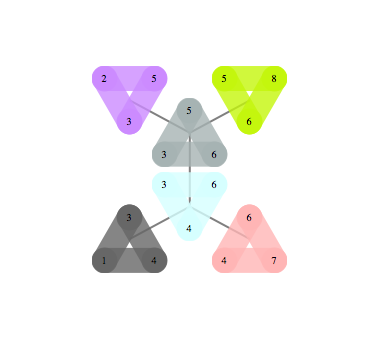 |
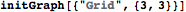
 |  |
100
010
001
111


words = Tuples[{0, 1}, 4];
distMat = Outer[HammingDistance, words, words, 1];
g = AdjacencyGraph[Map[Boole[# == 1] &, distMat, {2}]];
verts = FindIndependentVertexSet[g];
code = words[[verts]] Notebook
Notebook

v = 18;
row[k_] := PadLeft[IntegerDigits[Floor[k/(v + 1)*10^v]], v];
mat = row /@ Range[v];
Total@mat
Total@Transpose@mat
Export["~/research/qr/hinges/anim.swf", images];